
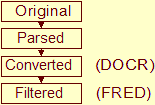
NMR Restraints Grid
 |
NMR Restraints Grid |

|
Result table
| image | mrblock_id | pdb_id | bmrb_id | cing | stage | position | program | type | subtype | subsubtype | item_count | other_prop |
|
|
20546 |
2g0q |
7007 | cing | 1-original | 0 | MR format | comment |
|
|
|
|
|
|
20547 |
2g0q |
7007 | cing | 1-original | 1 | XPLOR/CNS | distance | NOE | simple |
|
|
|
|
20548 |
2g0q |
7007 | cing | 1-original | 2 | XPLOR/CNS | dihedral angle |
|
|
|
|
|
|
20549 |
2g0q |
7007 | cing | 1-original | 3 | MR format | nomenclature mapping |
|
|
|
|
|
|
47054 |
2g0q |
7007 | cing | 2-parsed | 0 | STAR | comment |
|
|
0 |
|
|
|
47055 |
2g0q |
7007 | cing | 2-parsed | 0 | STAR | distance | NOE | simple | 1989 |
|
|
|
47056 |
2g0q |
7007 | cing | 2-parsed | 0 | STAR | dihedral angle |
|
|
189 |
|
|
|
47057 |
2g0q |
7007 | cing | 2-parsed | 0 | STAR | entry | full |
|
2178 |
|
|
|
424648 |
2g0q |
7007 | cing | 3-converted-DOCR | 0 | STAR | entry | full |
|
2178 |
|
|
|
424652 |
2g0q |
7007 | cing | 3-converted-DOCR | 0 | XML | entry | full |
|
|
|
|
|
424655 |
2g0q |
7007 | cing | 3-converted-DOCR | 0 | XPLOR/CNS | sequence |
|
|
|
|
|
|
424656 |
2g0q |
7007 | cing | 3-converted-DOCR | 0 | XPLOR/CNS | coordinate | ensemble |
|
|
|
|
|
424657 |
2g0q |
7007 | cing | 3-converted-DOCR | 0 | XPLOR/CNS | distance | general distance | ambi |
|
|
|
|
424658 |
2g0q |
7007 | cing | 3-converted-DOCR | 0 | XPLOR/CNS | dihedral angle |
|
|
|
|
|
|
424659 |
2g0q |
7007 | cing | 3-converted-DOCR | 0 | DYANA/DIANA | sequence |
|
|
|
|
|
|
424660 |
2g0q |
7007 | cing | 3-converted-DOCR | 0 | DYANA/DIANA | distance | general distance | ambi |
|
|
|
|
424661 |
2g0q |
7007 | cing | 3-converted-DOCR | 0 | DYANA/DIANA | distance | general distance | ambi |
|
LOWER_ONLY=true |
|
|
424663 |
2g0q |
7007 | cing | 3-converted-DOCR | 0 | DYANA/DIANA | dihedral angle |
|
|
|
|
|
|
424682 |
2g0q |
7007 | cing | 4-filtered-FRED | 0 | STAR | entry | full |
|
2109 |
|
|
|
424683 |
2g0q |
7007 | cing | 4-filtered-FRED | 0 | Wattos | check | stereo assignment | distance |
|
|
|
|
424684 |
2g0q |
7007 | cing | 4-filtered-FRED | 0 | Wattos | check | surplus | distance |
|
|
|
|
424685 |
2g0q |
7007 | cing | 4-filtered-FRED | 0 | Wattos | check | violation | distance |
|
|
|
|
424686 |
2g0q |
7007 | cing | 4-filtered-FRED | 0 | Wattos | check | violation | dihedral angle |
|
|
|
|
424687 |
2g0q |
7007 | cing | 4-filtered-FRED | 0 | Wattos | check | completeness | distance |
|
|
Contact the webmaster for help, if required. Saturday, April 20, 2024 9:33:20 AM GMT (wattos1)