
NMR Restraints Grid
 |
NMR Restraints Grid |
 |
Result table
| image | mrblock_id | pdb_id | cing | stage | position | program | type | subtype | subsubtype |
|
|
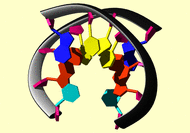
476131 |
1buf |
cing | 1-original | 3 | AMBER | distance | hydrogen bond | simple |
# r1=0, r4=r3+0.3; ref.: James (1994) Biochemistry 33, 354
# H-bond distance C1-G12, rk2, rk3 = 50% of kdist (10) = 5
&rst iat=17,365,-1,-1,iresid=0,r1=0,r2=2.81,r3=3.01,r4=3.31,
rk2=5,rk3=5,&end
&rst iat=20,366,r1=0,r2=2.85,r3=3.05,r4=3.35,&end
&rst iat=22,369,r1=0,r2=2.76,r3=2.96,r4=3.26,&end
# H-bond distance A2-T11
&rst iat=47,336,-1,-1,r1=0,r2=2.85,r3=3.05,r4=3.35,&end
&rst iat=50,337,-1,-1,r1=0,r2=2.72,r3=2.92,r4=3.22,&end
# H-bond distance A3-T10
&rst iat=79,304,-1,-1,r1=0,r2=2.85,r3=3.05,r4=3.35,&end
&rst iat=82,305,-1,-1,r1=0,r2=2.72,r3=2.92,r4=3.22,&end
# H-bond distance T4-A9
&rst iat=114,269,-1,-1,r1=0,r2=2.85,r3=3.05,r4=3.35,&end
&rst iat=115,272,-1,-1,r1=0,r2=2.72,r3=2.92,r4=3.22,&end
# H-bond distance T5-A8
&rst iat=146,237,-1,-1,r1=0,r2=2.85,r3=3.05,r4=3.35,&end
&rst iat=147,240,-1,-1,r1=0,r2=2.72,r3=2.92,r4=3.22,&end
# H-bond distance G6-C7
&rst iat=175,207,-1,-1,r1=0,r2=2.81,r3=3.01,r4=3.31,&end
&rst iat=176,210,-1,-1,r1=0,r2=2.85,r3=3.05,r4=3.35,&end
&rst iat=179,212,-1,-1,r1=0,r2=2.76,r3=2.96,r4=3.26,&end
Contact the webmaster for help, if required. Saturday, April 20, 2024 11:13:28 AM GMT (wattos1)