
NMR Restraints Grid
 |
NMR Restraints Grid |
 |
Result table
| image | mrblock_id | pdb_id | cing | stage | program | type |
|
|
32572 |
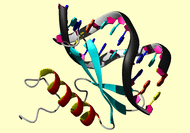
1gcc |
cing | 2-parsed | STAR | comment |
data_1gcc_MR_file_constraints
save_Conversion_project
_Study_list.Sf_category study_list
_Study_list.Entry_ID parsed_1gcc
_Study_list.ID 1
loop_
_Study.ID
_Study.Name
_Study.Type
_Study.Details
_Study.Entry_ID
_Study.Study_list_ID
1 "Conversion project" NMR . parsed_1gcc 1
stop_
save_
save_entry_information
_Entry.Sf_category entry_information
_Entry.ID parsed_1gcc
_Entry.Title "Original constraint list(s)"
_Entry.Version_type original
_Entry.Submission_date .
_Entry.Accession_date .
_Entry.Last_release_date .
_Entry.Original_release_date .
_Entry.Origination .
_Entry.NMR_STAR_version 3.1
_Entry.Original_NMR_STAR_version .
_Entry.Experimental_method NMR
_Entry.Experimental_method_subtype .
loop_
_Related_entries.Database_name
_Related_entries.Database_accession_code
_Related_entries.Relationship
_Related_entries.Entry_ID
PDB 1gcc "Master copy" parsed_1gcc
stop_
save_
save_global_Org_file_characteristics
_Constraint_stat_list.Sf_category constraint_statistics
_Constraint_stat_list.Entry_ID parsed_1gcc
_Constraint_stat_list.ID 1
loop_
_Constraint_file.ID
_Constraint_file.Constraint_filename
_Constraint_file.Software_ID
_Constraint_file.Software_label
_Constraint_file.Software_name
_Constraint_file.Block_ID
_Constraint_file.Constraint_type
_Constraint_file.Constraint_subtype
_Constraint_file.Constraint_subsubtype
_Constraint_file.Constraint_number
_Constraint_file.Entry_ID
_Constraint_file.Constraint_stat_list_ID
1 1gcc.mr . . "MR format" 1 comment "Not applicable" "Not applicable" 0 parsed_1gcc 1
1 1gcc.mr . . n/a 2 comment "Not applicable" "Not applicable" 0 parsed_1gcc 1
1 1gcc.mr . . XPLOR/CNS 3 distance NOE simple 0 parsed_1gcc 1
1 1gcc.mr . . n/a 4 comment "Not applicable" "Not applicable" 0 parsed_1gcc 1
1 1gcc.mr . . XPLOR/CNS 5 distance NOE simple 0 parsed_1gcc 1
1 1gcc.mr . . n/a 6 comment "Not applicable" "Not applicable" 0 parsed_1gcc 1
1 1gcc.mr . . XPLOR/CNS 7 distance NOE simple 0 parsed_1gcc 1
1 1gcc.mr . . n/a 8 comment "Not applicable" "Not applicable" 0 parsed_1gcc 1
1 1gcc.mr . . XPLOR/CNS 9 distance "hydrogen bond" simple 0 parsed_1gcc 1
1 1gcc.mr . . n/a 10 comment "Not applicable" "Not applicable" 0 parsed_1gcc 1
1 1gcc.mr . . XPLOR/CNS 11 distance "hydrogen bond" simple 0 parsed_1gcc 1
1 1gcc.mr . . n/a 12 comment "Not applicable" "Not applicable" 0 parsed_1gcc 1
1 1gcc.mr . . XPLOR/CNS 13 "dihedral angle" "Not applicable" "Not applicable" 0 parsed_1gcc 1
1 1gcc.mr . . n/a 14 comment "Not applicable" "Not applicable" 0 parsed_1gcc 1
1 1gcc.mr . . XPLOR/CNS 15 "dihedral angle" "Not applicable" "Not applicable" 0 parsed_1gcc 1
1 1gcc.mr . . "MR format" 16 "nomenclature mapping" "Not applicable" "Not applicable" 0 parsed_1gcc 1
stop_
save_
save_MR_file_comment_2
_Org_constr_file_comment.Sf_category org_constr_file_comment
_Org_constr_file_comment.Entry_ID parsed_1gcc
_Org_constr_file_comment.ID 1
_Org_constr_file_comment.Constraint_file_ID 1
_Org_constr_file_comment.Block_ID 2
_Org_constr_file_comment.Details "Generated by Wattos"
_Org_constr_file_comment.Comment
;
This file consists of A. NOE interproton distance restraints within protein,
B. those within DNA, C. intermolecular NOE restraints, D. hydrogen bond
restraints in protein, and E. theoretical restraints for DNA (see Omichinski
et al. Science 261, 438-446, 1993; Werner et al. Cell 81, 705-714, 1995 for
the DNA theoretical restraints).
All were used for the simulated anealing by XPLOR 3.1 but only A, B, and C
were used in the restraind molecular dynamical calculation by AMBER 4.1.
NOE restraints from bases 1, 2, 25, and 26, either, were not used in the
restrained molecular dynamics.
A. NOE interproton distance restraints within protein
;
save_
Contact the webmaster for help, if required. Thursday, April 18, 2024 5:49:15 PM GMT (wattos1)